PacBio and Loop-Seq sequencers, and newer Illumina sequencing platforms all output binned quality scores by default. Rather than a continuous distribution of quality scores, for example from 1 to 40 for the MiSeq platform, there are only three or four values. This can cause problems with processing the sequences, especially with the DADA2 algorithm for correcting and clustering sequences into ASVs.
If there are many different characters in the last line for every fastq sequence as in the image below, then the Q-scores are not binned. This is an example of sequeces produced by the MiSeq platform and the letters encode Q-values from 1 to 40.
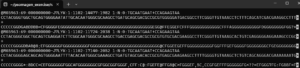
The image below shows sequences from the NovoSeq 6000 platform. There are only 3 characters in the quality score lines: comma, semicolon and upper case F. These encode the values 11, 25 and 37. If there were uncalled bases, the position would be assigned a 2.
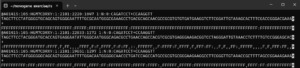
To decode the quality scores, look up the integer for the ASCII character and substract 33. You will need this information to process the sequences with the R version of DADA2. See the Processing Novogene Sequences exercise. There is currently no way of processing sequences with binned quality scores with the QIIME2 version (2026.1) of the DADA2 plugin.
